## Carbapenemase vs. ESBL: The Defining Distinction **Key Point:** Carbapenemase-producing organisms are defined by their ability to **hydrolyze carbapenems**, a property that distinguishes them from ESBL producers, which remain carbapenem-susceptible. ### Mechanism and Resistance Pattern Carbapenemases are β-lactamases (serine or metallo-enzymes) that have evolved the ability to cleave the β-lactam ring of carbapenems—the most stable β-lactams available. Unlike ESBL enzymes: - Carbapenemases confer resistance to **all β-lactams** including imipenem, meropenem, and ertapenem - This resistance is **NOT inhibited by clavulanic acid** because carbapenemases have different active site architecture - The resistance pattern is intrinsic to the enzyme's substrate specificity ### Comparative Resistance Profiles | Characteristic | ESBL | Carbapenemase | |---|---|---| | **Cephalosporin resistance** | 3rd gen (ceftriaxone, cefotaxime) | All generations | | **Carbapenem resistance** | ✗ Susceptible | ✓ Resistant | | **Clavulanic acid inhibition** | ✓ YES | ✗ NO | | **Clinical consequence** | Carbapenems effective | Carbapenems ineffective | | **Enzyme examples** | CTX-M, TEM, SHV | KPC, NDM, VIM, OXA-48 | **High-Yield:** The **Hodge test** or **Modified Carbapenem Inactivation Method (mCIM)** are rapid bedside tests to detect carbapenemase production. A positive test indicates carbapenem hydrolysis. **Mnemonic:** **KPC-NDM-VIM** = Major carbapenemases (Klebsiella Pneumoniae Carbapenemase, New Delhi Metallo-β-lactamase, Verona Integron-encoded Metallo-β-lactamase). ### Clinical Significance In the case presented, the isolate is resistant to imipenem and meropenem (carbapenems) but susceptible to colistin and tigecycline. This pattern is pathognomonic for carbapenemase production: - ESBL producers would be susceptible to carbapenems - The lack of clavulanic acid reversal confirms this is not an ESBL with AmpC co-production - Treatment options are severely limited (colistin, tigecycline, or combination therapy) **Clinical Pearl:** Carbapenem-resistant Enterobacteriaceae (CRE) are a major nosocomial threat. Early detection via carbapenemase screening is critical for infection control and therapeutic decision-making. [cite:Harrison 21e Ch 139; Park 26e Ch 3] 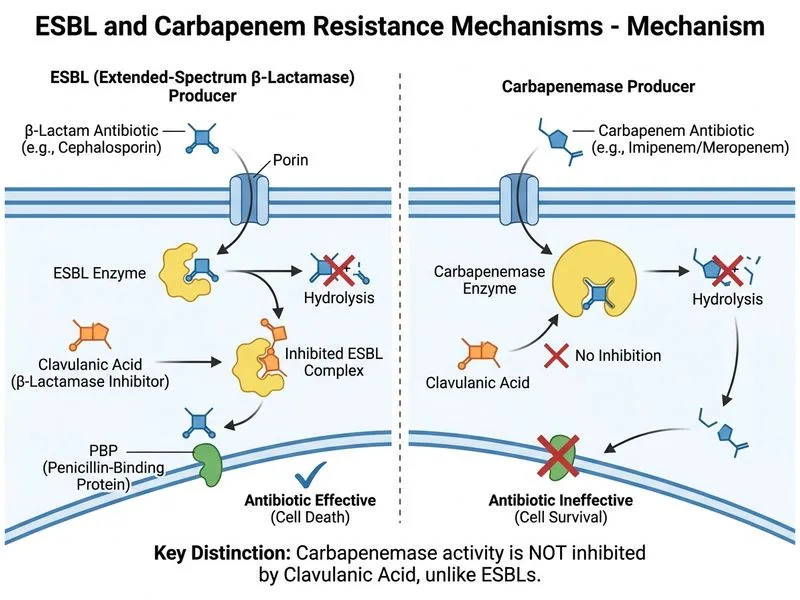
Sign up free to access AI-powered MCQ practice with detailed explanations and adaptive learning.